How structured AI pipelines can turn ambiguous clinical requests into reproducible workflows
Apr 29, 2026

Clinical data science workflows often begin with requests that appear simple on the surface. But in practice, these requests are rarely structured, and the underlying data is even less so. Roughly 80% of healthcare data exists in unstructured form, from clinician notes to wearable signals to survey responses, and the requests that emerge from that environment reflect the same messiness. A clinician or researcher might ask: can we predict next-day pain spikes from this wearable data? Behind this question are multiple implicit decisions, including target definitions, time windows, leakage constraints, feature construction, and evaluation metrics, that must be made explicit before any modeling can begin. In most workflows today, this translation from intent to task is manual, iterative, and error-prone.
This ambiguity is not just inconvenient. It is a primary source of failure in clinical AI. Most failures do not come from model performance, but from mis-specified tasks. Common issues include target leakage (using future data unintentionally), inconsistent definitions across sites, misaligned evaluation metrics, and hidden assumptions in preprocessing. Manual medical record abstraction has a pooled error rate of approximately 7%, which is large enough to compromise the statistical power of clinical trials. Even that figure understates the variability introduced when humans interpret ambiguous clinical intent.
Why Clinical Intent is Hard to Parse
Clinical and research intent is inherently complex. Key details are often missing or assumed, definitions like “pain spike” may vary across studies, and requests frequently mix high-level goals with low-level constraints. These are not edge cases: in one analysis of 334 pain trials, researchers identified 50 different outcome measures for pain, with only 14% adhering to a standardized core set. Inputs also span multiple modalities, including EHRs, wearables, and surveys, further complicating how intent is interpreted. Human speech compounds this challenge, introducing disfluencies, informal phrasing, and implicit assumptions. For this reason, we used voice as a deliberate stress test: a system that can resolve intent from dictated requests can handle the full range of ambiguous clinical input.
From Complex Intent to Structured Task: The Intent Clarifier
We developed an intent clarifier, an automated system that interprets ambiguous clinical requests and translates them into fully specified modeling tasks. This was developed as part of a broader end-to-end pipeline that converts unstructured, complex requests into reproducible modeling workflows.
At a high level, the system:
Accepts input in any form, including voice, transcribed via MedASR (optimized for medical dictation)
Extracts structured intent from the input
Identifies missing or ambiguous fields
Asks targeted clarification questions as needed
Outputs a fully specified modeling task
Rather than requiring users to provide perfectly structured inputs upfront, the system iteratively resolves ambiguity, similar to how a collaborator would. The core challenge is not transcription, it is clarification. The real problem is that clinical intent arrives incomplete: objectives are vague, target variables are undefined, and assumptions go unstated. The system first attempts to infer the objective, target variable, available datasets, and initial assumptions, then identifies what is missing and asks only the minimum number of targeted questions needed to complete the task definition.
By enforcing structure before modeling begins (for example, standardizing how targets are defined, resolving missing parameters such as prediction windows or evaluation metrics, and identifying potential leakage risks in feature construction), the system reduces iteration cycles between clinicians and data scientists, prevents silent errors that invalidate results, and enables consistent workflows across datasets and institutions. This shifts clinical AI development from ad hoc and reactive workflows to structured and reproducible ones.
Case Study: Pain Spike Prediction
We evaluated this approach using real-world data from patients with chronic pain and opioid use disorder, collected in collaboration with the APT Foundation. This setting is particularly well-suited for testing intent clarification, as defining a task like “predict next-day pain spikes” requires making multiple implicit decisions around target definitions, time windows, feature construction, and leakage constraints. The dataset includes wearable signals such as sleep, activity, and heart rate proxies, along with patient-reported surveys capturing pain, stress, and mental health, resulting in approximately 180 derived daily features across 25 patients.
The modeling task was to predict next-day evening pain spikes, defined as values above the 70th percentile of self-reported pain, based on clinician-informed thresholds. While prior work focused on model performance, this experiment focused on a different question: can we reliably translate ambiguous clinical intent into a valid, reproducible modeling pipeline?
Once intent is clarified, the system generates:
A structured task contract (objective, target, metrics, leakage constraints)
A reproducible code bundle (train.py, config, requirements)
A modeling plan (time-series splits, feature groups, baseline model)
Executed output, including baseline model results and evaluation metrics
This ensures that the downstream workflow is deterministic (no hidden assumptions), reproducible across environments, and auditable for clinical and research settings.
How It Works in Practice
Here is how the full pipeline works end to end:
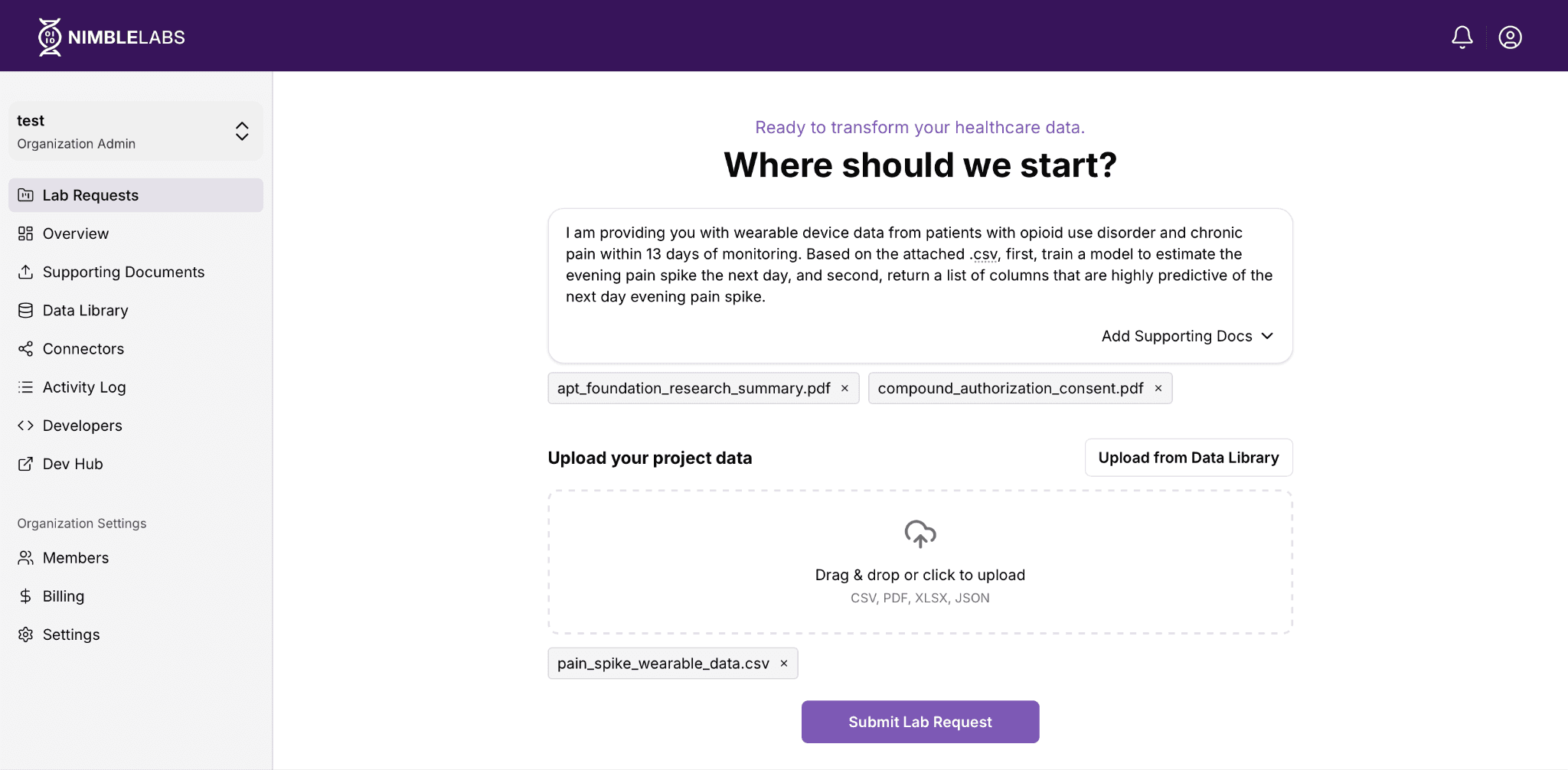
Step 1: Submit your request. A researcher navigates to NimbleLabs (Nimblemind’s user-facing platform for submitting data requests and managing datasets) and completes the following steps:
Describe the task. The user describes their goal in plain language (no structured query required).
Provide input via voice. In our case study, the user provided the following request via voice, which was transcribed into text for processing:
“I am providing you with wearable device data from patients with opioid use disorder and chronic pain within 13 days of monitoring. Based on the attached .csv, first, train a model to estimate the evening pain spike the next day, and second, return a list of columns that are highly predictive of the next day evening pain spike.”
Upload data and context. The user uploads their dataset and any supporting documents, such as research consent forms or study summaries.
Step 2: Request enters the pipeline. Once submitted, the request is first processed by the intent clarifier, which extracts the core task, identifies missing or ambiguous fields, and resolves them through structured interpretation. The system then logs the fully specified task, queues it for execution, and provides an estimated turnaround.
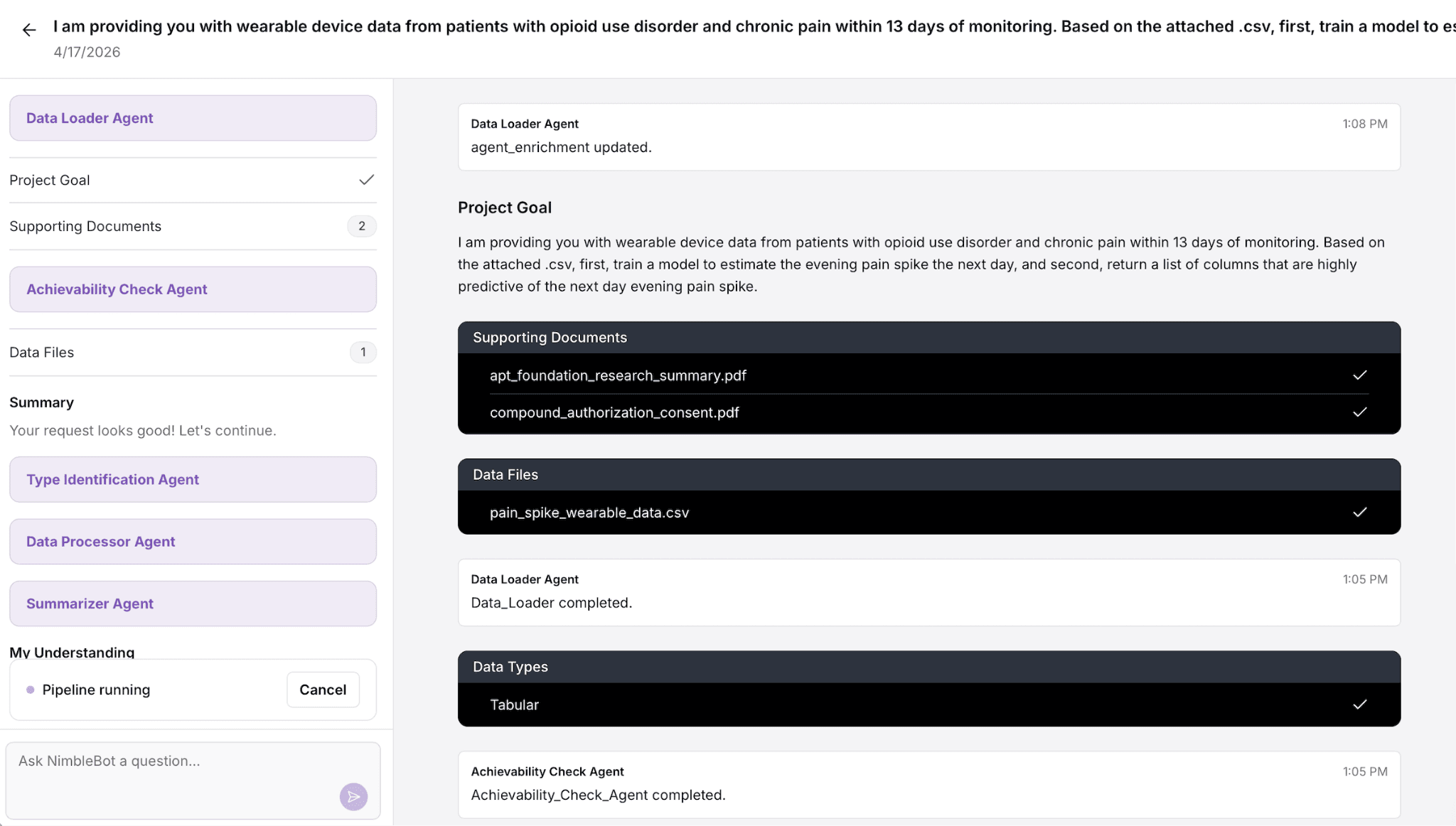
Step 3: Confirmation and transparency. The user receives an email confirmation summarizing exactly what was submitted and inferred: datasets, supporting documents, the structured task definition, and the resolved intent. This creates an auditable record of what was asked and how it was interpreted before any modeling begins.
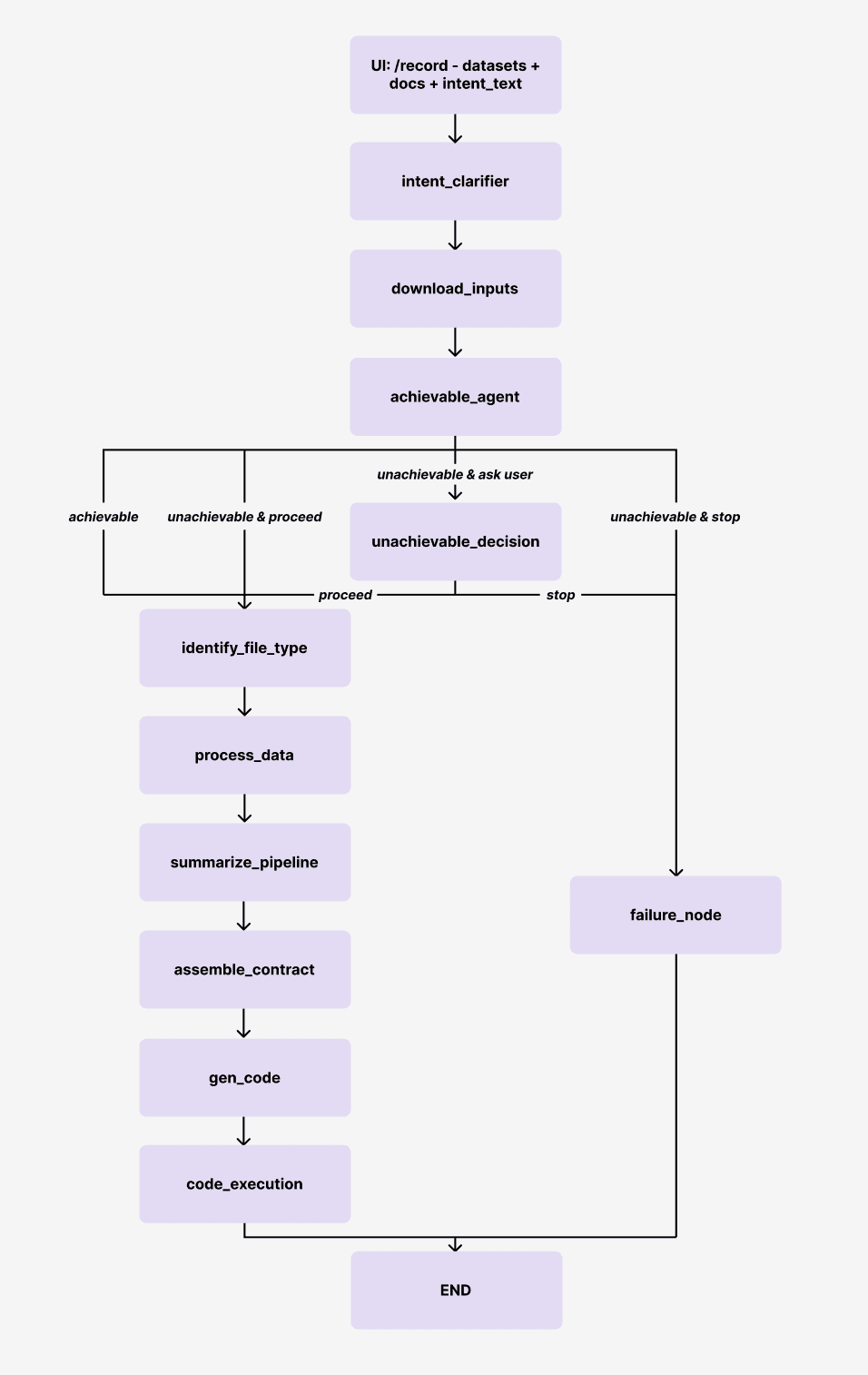
Step 4. The pipeline resolves intent and executes. Behind the scenes, the system runs through a structured sequence of steps. The pipeline graph below illustrates the full flow:
Intent clarification. The request is interpreted by the intent_clarifier, which extracts the task and resolves missing or ambiguous fields through structured clarification.
Validation. The clarified intent is validated by an intent_guard to ensure consistency and completeness.
Feasibility assessment. The achievable_agent determines whether the request can be executed with the available data and constraints.
Data understanding and processing. The system identifies file types, parses inputs, and prepares the data for modeling.
Task structuring. A structured task contract is assembled, and the pipeline is summarized.
Code generation and execution. The system generates code, runs the workflow, and produces results.
If the request is unachievable, the system either stops or surfaces a decision point for the user, rather than silently failing or producing invalid results.
The user can view the full request details inside the platform, including the structured intent, uploaded files, and result outputs, at any stage of processing.
Looking Ahead
At Nimblemind, this approach reflects a broader focus on building systems that make clinical AI workflows traceable (every assumption is explicit), reproducible (same input generates same output), and deployable across real-world environments. The intent clarifier is a foundational layer in this stack, ensuring that before any model is trained, the system and the user are aligned on what is actually being asked.
As clinical AI systems begin to ingest more diverse forms of data (including text, structured records, wearable signals, and voice), the way users specify tasks is also evolving. Instead of filling out rigid forms with predefined fields, users increasingly describe their goals in natural language, often with incomplete or ambiguous inputs. In that setting, the challenge is not just understanding language, but resolving intent into a well-defined, executable task.
This work shows that:
Clinically complex workflows are feasible
Intent can be systematically clarified
Structured pipelines can be generated from unstructured input
Future work will extend this to more complex multimodal tasks, iterative workflows, and tighter integration with deployment and monitoring systems.
Try the system → https://nimblelabs.ai/agent
Read the underlying study → https://arxiv.org/abs/2511.19577